- The good news is that we all basically agreed on the viral sequences.
- The bad news is that we couldn't agree on about 10% of the bacterial sequences.
- The challenging news is that there wasn't enough information to decide one way or another for more than 30% of the bacterial sequences and more than 80% of the fungal sequences.
Jake Beal's Next Step
Irregular thoughts and discussion about my life as a scientist.
Monday, May 06, 2024
Does your biosecurity screening work?
Tuesday, November 14, 2023
How do you describe genetic construction plans?
Monday, April 03, 2023
BLAST vs. custom tools for pathogen identification
Our analysis of issues with using BLAST vs. NCBI for pathogen identification is out today: “Studying pathogens degrades BLAST-based pathogen identification.” This paper is the full published version of the preprint I posted about a few months ago, investigating an emergent dynamic, in which biological research and development ends up contaminating public databases with chimeric material that can confound biosecurity systems that trust those databases.
The most important addition between the preprint and this final version was to make direct head-to-head comparisons between BLAST vs. NCBI and two tools specifically designed for biosecurity analysis, our own FAST-NA Scanner and a free tool called SeqScreen (there are other tools we'd like to have compared with as well, but they were not available for comparison).
As predicted, the actual biosecurity tools completely dominated over BLAST vs. NCBI, making more than an order of magnitude less mistakes---not a surprise, but nice to see experimentally validated. In fact, each biosecurity tool only made one mistake in judgement, and in both cases it was the same mistake that NCBI did, which is an important lesson: the big NCBI databases aren't bad, they're just dirty, and so they just need a lot of care and refinement when they're being put to a use (like biosecurity determinations) where mistakes can be costly and dangerous.
This is important for biosecurity, but I also think people need to be aware of this in the larger scientific world as well. In biology, curation quality really matters, and many people are far too blasé about the potential impact of dirty data on their applications. If you want to do biosecurity right, you need to use an actual biosecurity tool and not just trust the databases. I'm sure the same applies for many aspects of medicine, diagnostics, etc., and I fear that not enough people are taking these issues seriously.
Wednesday, July 27, 2022
Multicolor Plate Reader Fluorescence Calibration
Just out in OUP Synthetic Biology, "Multicolor Plate Reader Fluorescence Calibration" extends our prior work on calibrating green fluorescence and cell count to calibrate red and blue fluorescence as well. The results are no surprise (if we can use a green dye, we ought to be able to use other dyes too), but it's valuable to have specific recommendations for dyes to use and to have an interlab study validate that yes, they really do perform as well as the others.
So everybody out there listening, please start using sulforhodamine-101 to calibrate your red fluorescence and Cascade Blue to calibrate your blue fluorescence! Everybody who uses your data will thank you for providing equivalent molecule/cell estimates rather than irreproductible arbitrary or relative units.
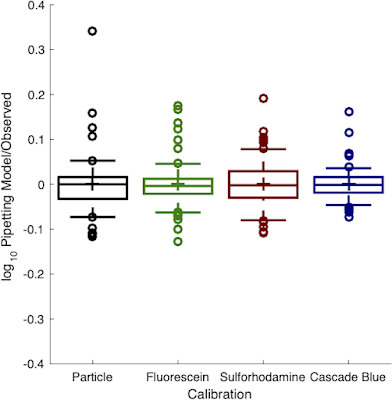 |
| Red and blue fluorescence calibrants were just as precise as the prior green and cell-count calibrants |
The paper also reports on some of the travails we ran into making the study work: some of the fluorescent proteins we wanted to try out didn't work in our hands, and there were miscellaneous other problems: a promoter sequence got messed up, some things wouldn't synthesize, one of the plasmids seemed problematic, and timing problems meant not all labs could run all constructs.
Problems like that are frustrating, but ultimately I'm happier reporting them than burying them. Remember: if you read a synthetic biology study with lab work and it doesn't talk about failures, it just means they either aren't aware of them or else they've pruned them from the narrative! Calibration methods like these help us see better when things go wrong and understand what's happened.
Thursday, July 14, 2022
Studying Pathogens Degrades BLAST-based Pathogen Identification
Using the BLAST algorithm to search the NCBI databases is the typical way one goes about identifying a DNA sequence, so it's been the typical way biosecurity systems decide if something is potentially a dangerous pathogen or toxin too. Problem is, that's not what BLAST and those databases were designed for, and we've observed that they aren't working as well for that purpose as they used to, as we report in our new preprint: "Studying Pathogens Degrades BLAST-based Pathogen Identification"
Specifically, we've found an inherent problem that is growing in seriousness due to a non-obvious emergent dynamic. Now that sequencing and bioengineering tools are getting much more accessible, lots of sequences are being studied by modifying them with "tool" sequences like purification tags, fluorescent proteins, stabilizing sequences, etc. Those sequences get (appropriately) classified based on what's being studied, and now you've got chimeric material that includes both the subject of study and the bioengineering tool. Then when you run BLAST on a sequence with that tool, you start finding that tools are classified as what they're used to study.
This doesn't seem to be much of a problem for most uses of BLAST against NCBI, but it's poisonous for making biosecurity decisions, since it can cause benign sequences to be classified as dangerous or vice versa. Moreover, the effect gets stronger the more problematic a pathogen is (since more sequences are recorded) and the more useful a tool is (since more chimeric material is produced), meaning that the problem is most likely to occur in the most important. For example, over the last two years, quite a lot of stuff has started coming back as COVID-19, since everybody in the world is studying COVID-19 with all of the tools that they can get their hands on.
This is a serious problem, and it's not likely to get better, since NCBI and BLAST aren't doing the wrong thing: they're just getting less suitable to use as a short-cut for doing something that they were never designed to do.
So how do we fix it? Switch to tools that are actually designed for pathogen identification. We've got one (FAST-NA Scanner), and a whole bunch of other folks worked on the same problem in the FunGCAT program. The solutions are there, we just have to help folks switch to them.
Wednesday, July 13, 2022
pySBOL3: SBOL3 for Python Programmers
Tuesday, July 05, 2022
Functional Synthetic Biology
Synthetic biology isn't about sequences. Don't agree? Tell me what this is without looking it up: atgcgtaaaggagaagaacttttcactggagttgtcccaattcttgttga
Tell you what, I'll give you a hint, make it easy. It's a coding sequence translating to MRKGEELFTGVVPILV. Everybody knows this one, right?
How about this instead?
Don't get me wrong, sequences are important. But right now we're living with a mis-match in synthetic biology, where most of our discussions about design are about function, but nearly all of our tooling is heavily focused on sequences (e.g., GenBank format), with any information about function tacked on as an afterthought or else confined to specialized databases that each pose their own sui generis integration problem.
We need a new focus on functional synthetic biology, and that's one of the things we've been working on in the iGEM Engineering Committee. We're trying to change how we do synthetic biology, so that we can pull together the work that lots of people have been doing on calibration, insulation, characterization, context effects, modeling, assembly, etc., in one place and make at least a small class of synthetic biology engineering really simple and predictable.
We aren't there yet, but we've gotten to the point where we think we've figured out some of the important shifts in thinking, representation, and tooling that need to happen in order to make functional synthetic biology possible. If you're interested in this too, I encourage you to read more in our newly available pre-print on Functional Synthetic Biology.
Thursday, May 05, 2022
AI for Synthetic Biology
Several of my colleagues have been organizing an series of "AI for SynBio" workshops over the last few years. I've been to some and they have been both stimulating and enjoyable. Now they have an article out in Communications of the ACM, along with a nice short video in which Aaron Adler introduces this increasingly important cross-disciplinary interaction for folks who aren't familiar with one or both of the subjects.
Friday, April 22, 2022
Talking measurement and standards with "The Living Revolution"
Yesterday I had an enjoyable conversation with Luke Roche and Sara Knurowska, who do a podcast called "The Living Revolution." They'd read some of my work on measurement, which led inevitably to a wide-ranging discussion including fundamental principles in engineering and science, when to standardize (or not), SBOL, etc.
Check out the podcast here (if it works for in your browser), or on Spotify or Apple Podcasts
Monday, February 14, 2022
"Meeting Measurement Precision" published in ACS Synthetic Biology
The pre-print that I wrote about in October, "Meeting Measurement Precision Requirements for Effective Engineering of Genetic Regulatory Networks", has just been published in ACS Synthetic Biology. Check out the official final version at: https://doi.org/10.1021/acssynbio.1c00488!





